Overview of the Python HPC landscape 🚀¶



Python for High-Performance Computing?¶
Fast prototyping (Numpy!)
Popular:
Well-known
Several great libraries
Share ideas between developers / scientists
Popularity counts
Readability counts
Expressivity counts
Anyway, one needs a good and well-known scripting language so yes!
(even considering Julia)
Where / when should we stop ?¶
Python: a programming language, compromises ⚖️¶
Designed for fast prototyping & "glue" codes together
Generalist + easy to learn ⇒ huge and diverse community 👨🏿🎓🕵🏼 👩🏼🎓 👩🏽🏫👨🏽💻👩🏾🔬 🎅🏼 🌎 🌍 🌏
Expressivity and readability
Not oriented towards high performance
(fast and easy dev, easy debug, correctness)
Highly dynamic 🐒 + introspection (
inspect.stack())Automatic memory management 💾
All objects encapsulated 🥥 (PyObject, C struct)
Objects accessible through "references" ➡️
Usually interpreted
Python interpreters¶
CPython 
Interpreted (nearly) instruction per instruction, (nearly) no code optimization
The numerical stack (Numpy, Scipy, Scikits, ...) based on the CPython C API (CPython implementation details)!
PyPy 
Optimized implementation with tracing Just-In-Time compilation
The CPython C API is an issue! PyPy can't accelerate Numpy code!
Micropython 
For microcontrollers
mylist = [1, 3, 5]
list: array of references towards PyObjects
The C / Python border¶
arr = 2 * np.arange(10)
print(arr[2])
4
Pure Python terrible 🐢 (except with PyPy)...
from math import sqrt
my_const = 10.
result = [elem * sqrt(my_const * 2 * elem**2) for elem in range(1000)]
but even this is not very efficient (temporary objects)...
import numpy as np
a = np.arange(1000)
result = a * np.sqrt(my_const * 2 * a**2)
Even slightly worth with PyPy 🙁
Is Python efficient enough?¶
Book¶

Performance (generalities)¶
Measure ⏱, don't guess! Profile to find the bottlenecks.¶
Cprofile (pstats, SnakeViz), line-profiler, perf, perf_events
Do not optimize everything!¶
"Premature optimization is the root of all evil" (Donald Knuth)
80 / 20 rule, efficiency important for expensive things and NOT for small things
CPU or IO bounded problems¶
Use the right algorithms and the right data structures!¶
For example, using Numpy arrays instead of Python lists...
Unittests before optimizing to maintain correctness!¶
unittest, pytest
"Crunching numbers" and computers architectures¶
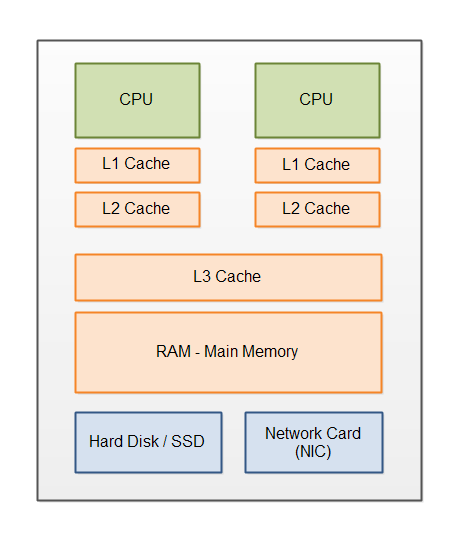
CPU optimizations¶
pipelining, hyper-threading, vectorization, advanced instructions (simd), ...
important to get data aligned in memory (arrays)
Proper compilation needed for high efficiency !¶
Compilation to virtual machine instructions¶
What does CPython (compile, "byte code", nearly no optimization, see dis module)
Compilation to machine instructions¶
Just-in-time
Has to be fast (warm up), can be hardware specific
Ahead-of-time
Can be slow, hardware specific or more general to distribute binaries
Compilers are usually good for optimizations! Better than most humans...
Transpilation¶
From one language to another language (for example Python to C++)
Different parallel strategies¶

Different parallel strategies¶
One process split in light subprocesses called threads 👩🏼🔧 👨🏼🔧👩🏼🔧 👨🏼🔧¶
handled by the OS
share memory and can use at the same time different CPU cores
How?
OpenMP (Natively in C / C++ / Fortran. For Python: Pythran, Cython, ...)
In Python:
threadingandconcurrent.futures
⚠️ in Python, one interpreter per process (~) and the Global Interpreter Lock (GIL)...
In a Python program, different threads can run at the same time (and take advantage of multicore)
But... the Python interpreter runs the Python bytecodes sequentially !
Terrible 🐌 for CPU bounded if the Python interpreter is used a lot !
No problem for IO bounded !
2 other packages for parallel computing with Python¶
- dask
- joblib
Python for HPC: first a glue language¶
Many tools to interact with static languages:
ctypes, cffi, cython, cppyy, pybind11, f2py, pyo3, ...
Glue together pieces of native code (C, Fortran, C++, Rust, ...) with a nice syntax
⇒ Numpy, Scipy, ...
Remarks:
Numpy: great syntax for expressing algorithms, (nearly) as much information as in Fortran
Performance of
a @ b(Numpy) versusa * b(Julia)?
Same! The same library is called! (often OpenBlas or MKL)
General principle for perf with Python (not fully valid for PyPy):¶
Don't use too often the Python interpreter (and small Python objects) for computationally demanding tasks.
Pure Python
→ Numpy
→ Numpy without too many loops (vectorized)
→ C extensions
But ⚠️ ⚠️ ⚠️ writting a C extension by hand is not a good idea ! ⚠️ ⚠️ ⚠️
No need to quit the Python language to avoid using too much the Python interpreter !¶
Tools to¶
compile Python
write C extensions without writing C
Cython, Numba, Pythran, Transonic, PyTorch, ...
First conclusions¶
- Python great language & ecosystem for sciences & data
Performance issues, especially for crunching numbers 🔢
⇒ need to accelerate the "numerical kernels"
Many good accelerators and compilers for Python-Numpy code
- All have pros and cons!
⇒ We shouldn't have to write specialized code for one accelerator!
Other languages don't replace Python for sciences
Modern C++ is great and very complementary 💑 with Python
Julia is interesting but not the heaven on earth