Python accelerators¶
General principle for perf with Python (not fully valid for PyPy):¶
Don't use too often the Python interpreter (and small Python objects) for computationally demanding tasks.
Pure Python
→ Numpy
→ Numpy without too many loops (vectorized)
→ C extensions
But ⚠️ ⚠️ ⚠️ writting a C extension by hand is not a good idea ! ⚠️ ⚠️ ⚠️
No need to quit the Python language to avoid using too much the Python interpreter !¶
Tools to¶
compile Python
write C extensions without writing C
Cython, Numba, Pythran, Transonic, ...

Langage: superset of Python
A great mix of Python / C / CPython C API!
Very powerfull but a tool for experts!
Easy to study where the interpreter is used (
cython --annotate).Very mature
Now able to use Pythran internally...
My experience: large Cython extensions difficult to maintain
Numba: (per-method) JIT for Python-Numpy code¶

- Very simple to use (just add few decorators) 🙂
from numba import jit
@jit
def myfunc(x):
return x**2
"nopython" mode (fast and no GIL) 🙂
Also a "python" mode 🙂
GPU and Cupy 😀
Methods (of classes) 🙂
Python decorators¶
def mydecorator(func):
# do something with the function
print(func)
# return a(nother) function
return func
@mydecorator
def myfunc(x):
return x**2
<function myfunc at 0x7eff33735820>
This mysterious syntax with @ is just syntaxic sugar for:
def myfunc(x):
return x**2
myfunc = mydecorator(myfunc)
<function myfunc at 0x7eff5d679790>
Concrete example: Fibonacci series¶
Definition:
$F_0 = 0$
$F_1 = 1$
$F_n = F_{n-1} + F_{n-2}$
First elements:
0, 1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89, 144, ...
def basic_fib(n):
# works but slow
if n < 2:
return n
else:
return basic_fib(n-1) + basic_fib(n-2)
# take some seconds
print(basic_fib(34))
5702887
Fibonacci series: keep calls in memory with a dict¶
Basic solution can only go up to n = 30 - 35 as the number of calls grows exponentially.
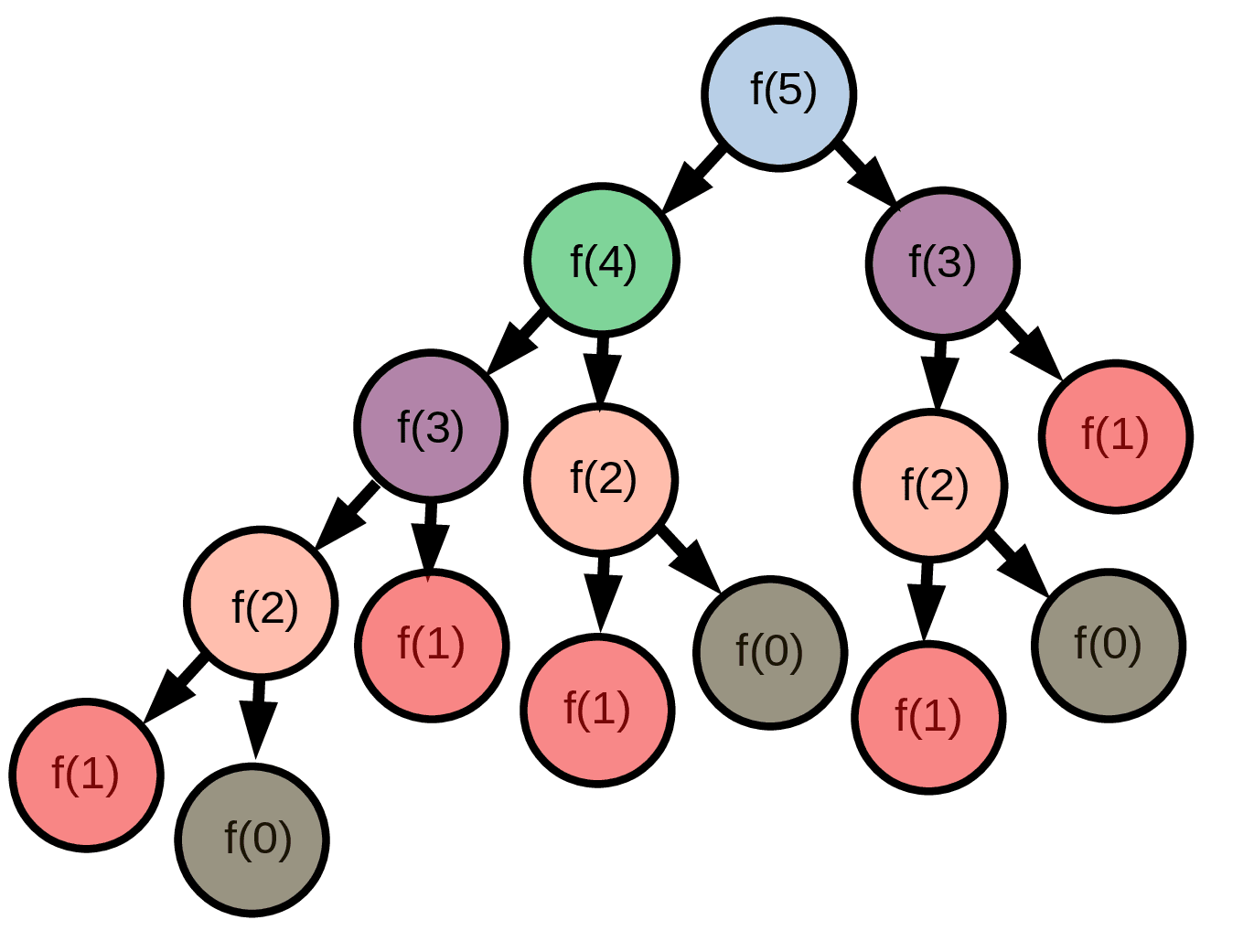
To overcome this limitations we will use a dictionnary that keeps the previous calls to the function in memory to avoid redundancy in function calls. This is called caching.
1: Write a function that takes an int and a dict as arguments. Use the dict to remove duplicate calls.
2: Write a general cache that takes a function as an argument and does the caching. You can then pass basic_fib as an argument and get a cached function.
3: The general cache is a decorator, use it with the @ syntax.
def basic_fib(n):
# reference function
if n < 2:
return n
else:
return basic_fib(n-1) + basic_fib(n-2)
# Step 1
def fib_with_dict(n, cache={}): # now takes a dict as an argument
# write the function body
return None
# bigger values should be accessible now:
print(fib_with_dict(100))
None
# Step 2
def general_cache(func): # take a function as an argument
# write the body of the function, do not implement the Fibonacci series here
# returns a function that behaves as the original function
# except that it checks whether the result is already stored in a dict
return cached_func
# Step 3, profit! Nothing to do, your general_cache should work as a decorator
# uncomment when general_cache is ready
# @general_cache
def basic_fib_decorated(n):
if n < 2:
return n
else:
return basic_fib_decorated(n-1) + basic_fib_decorated(n-2)
Solution step 1: use a dict as an argument¶
# one possible solution
def fib_with_dict(n, cache={}):
if n in cache:
return cache[n]
else:
if n < 2:
return n
else:
cache[n] = fib_with_dict(n-1, cache) + fib_with_dict(n-2, cache)
return cache[n]
print(fib_with_dict(100))
354224848179261915075
Solution step 2: a more general cache¶
def basic_fib(n):
# do not touch
if n < 2:
return n
else:
return basic_fib(n-1) + basic_fib(n-2)
def general_cache(func):
# Take a function as an argument
cache = {}
def cached_func(*args):
if args not in cache:
cache[args] = func(*args)
return cache[args]
# returns a function that behaves as the original function
# except that it checks whether the result is already stored in a dict
return cached_func
basic_fib = general_cache(basic_fib)
print(basic_fib(100))
354224848179261915075
With a standard library¶
from functools import lru_cache
@lru_cache(maxsize=1024)
def basic_fib_decorated(n):
if n < 2:
return n
else:
return fib(n-1) + fib(n-2)
Bonus question: Why is the first line cached but not the second one? (Try yourself)
basic_fib = general_cache(basic_fib)
basic_fib_cached = general_cache(basic_fib)
Numba: (per-method) JIT for Python-Numpy code¶

Sometimes not as much efficient as it could be 🙁
(usually slower than Pythran / Julia / C++)
- Only JIT 🙁
- Not good to optimize high-level NumPy code 🙁
Pythran: AOT compiler for module using Python-Numpy¶
Transpiles Python to efficient C++

Good to optimize high-level NumPy code 😎
Extensions never use the Python interpreter (pure C++ ⇒ no GIL) 🙂
Can produce C++ that can be used without Python
Usually very efficient (sometimes faster than Julia)
High and low level optimizations
(Python optimizations and C++ compilation)
SIMD 🤩 (with xsimd)
Understand OpenMP instructions 🤗 !
Used in Scipy: portable and stable
Can use and make PyCapsules (functions operating in the native word) 🙂
High level transformations¶
# calcul of range
print_optimized("""
def f(x):
y = 1 if x else 2
return y == 3
""")
def f(x):
return 0
# inlining
print_optimized("""
def foo(a):
return a + 1
def bar(b, c):
return foo(b), foo(2 * c)
""")
def foo(a):
return a + 1
def bar(b, c):
return ((b + 1), ((2 * c) + 1))
# unroll loops
print_optimized("""
def foo():
ret = 0
for i in range(1, 3):
for j in range(1, 4):
ret += i * j
return ret
""")
def foo():
ret = 0
ret += 1
ret += 2
ret += 3
ret += 2
ret += 4
ret += 6
return 18
# constant propagation
print_optimized("""
def fib(n):
return n if n< 2 else fib(n-1) + fib(n-2)
def bar():
return [fib(i) for i in [1, 2, 8, 20]]
""")
import functools as __pythran_import_functools
def fib(n):
return n if (n < 2) else (fib((n - 1)) + fib((n - 2)))
def bar():
return [1, 1, 21, 6765]
# advanced transformations
print_optimized("""
import numpy as np
def wsum(v, w, x, y, z):
return sum(np.array([v, w, x, y, z]) * (.1, .2, .3, .2, .1))
""")
import numpy as __pythran_import_numpy
def wsum(v, w, x, y, z):
return builtins.sum(((v * 0.1), (w * 0.2), (x * 0.3), (y * 0.2), (z * 0.1)))
Pythran: AOT compiler for module using Python-Numpy¶
- Compile only full modules (⇒ refactoring needed 🙁)
Only "nopython" mode
limited to a subset of Python
- only homogeneous list / dict 🤷♀️
- no methods (of classes) 😢 and user-defined class
limited to few extension packages (Numpy + bits of Scipy)
pythranized functions can't call Python functions
No JIT: need types (written manually in comments)
Lengthy ⌛️ and memory intensive compilations (especially with gcc, less with clang)
Debugging 🐜 Pythran requires C++ skills!
No GPU (maybe with OpenMP 4?)
 compilers unable to compile Pythran C++11 👎
compilers unable to compile Pythran C++11 👎
First conclusions¶
- Python great language & ecosystem for sciences & data
Performance issues, especially for crunching numbers 🔢
⇒ need to accelerate the "numerical kernels"
Many good accelerators and compilers for Python-Numpy code
- All have pros and cons!
⇒ We shouldn't have to write specialized code for one accelerator!
Keep your Python-Numpy code clean and "natural" 🧘
Clean type annotations (🐍 3)
Easily mix Python code and compiled functions
JIT based on AOT compilers
Methods (of classes) and blocks of code
Transonic: examples from real-life packages¶
JIT (
@jit)AOT compilation for functions and methods (
@boost)Blocks of code (with
if ts.is_transpiled:)Parallelism with a class (adapted from Olivier Borderies)
omp/tsp.py (OpenMP) and tsp_concurrent.py (concurrent - threads)
Also compatible with MPI!
Works also well in simple scripts and IPython / Jupyter.
# abstract syntax tree
import ast
tree = ast.parse("great_tool = 'Beniget'")
assign = tree.body[0]
print(f"{assign.value.s} is a {assign.targets[0].id}")
Beniget is a great_tool
Write the (Pythran) files when needed
Compile the (Pythran) files when needed
Use the fast solutions when available
Success: publication in Nature Astronomy¶
"Reducing the ecological impact of computing through education and Python compilers"
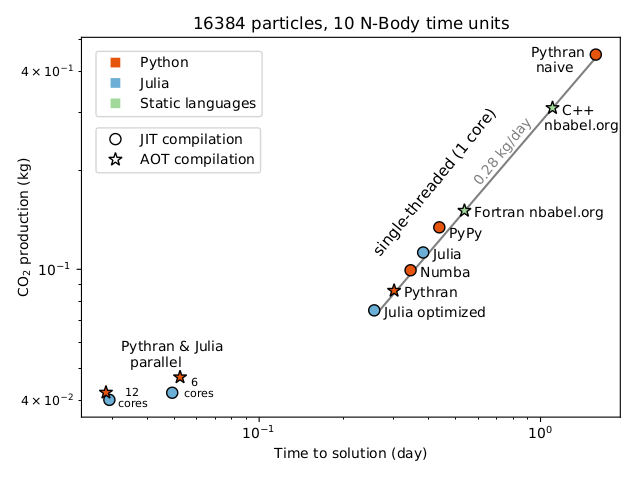
- Lien vers la note : https://rdcu.be/ciO0J
- Page sur le site du journal : https://www.nature.com/articles/s41550-021-01342-y
- Dépôt des benchmarks avec le code : https://github.com/paugier/nbabel
- Un article "grand public" sur ce travail : https://insis.cnrs.fr/fr/cnrsinfo/reduire-limpact-environnemental-du-calcul-scientifique-par-loptimisation-des-codes-et-la
PyTorch¶
Conclusions¶
Many good accelerators and compilers for Python-Numpy code
Very good results for "standard types" (arrays of numbers on CPU and GPU)
Limitation: high performance with user-defined numerical types